Mission of the Program in Cancer Biology
To train new leaders in the field of Cancer Biology that will develop new knowledge that will translate into improved detection, diagnosis, prognosis, prevention, or treatment of cancer.
Research Areas of Emphasis
- Cancer Immunity, host tumor interactions, and angiogenesis
- Cancer Precision Medicine—targeted therapies and drug resistance using mouse modeling, human tumor tissues, and systems approaches
- Bioinformatic analyses of tumor heterogeneity including genome, proteome, metabolome, and immunome components during tumor progression
- Basic Cancer Biology—tumor progression, invasion and metastasis
Steering Committee
Rachelle Johnson, Program Director and Director of Graduate Studies
Jin Chen
Barbara Fingleton
Jeffrey Rathmell
W. Kimryn Rathmell
Julie Rhoades
Ann Richmond
Alissa Weaver
Chris Williams
News & Events

Jared Rhodes
(Mentor James Goldenring, MD, PhD)
Jared won the best student graduate poster at the 23rd Annual Retreat for Cancer Biology! His research in Goldenring Lab examines fibroblasts and how they have recently been shown to promote the dysplastic transition of metaplastic lineages in the stomach. Specifically, conditioned media (CM) from gastric metaplasia and cancer-derived fibroblasts accelerated dysplastic progression in metaplastic gastroids. It remains unclear which factors secreted from fibroblasts cause this progression. Therefore, he performed scRNA-sequencing on normal, metaplasia, and cancer-derived fibroblasts. This revealed expression differences of many secreted factor encoding genes. Metaplastic gastroids were cultured in 30 and 50 kDa filtered CM from cancer-derived fibroblasts to narrow down candidate factors. 30 kDa filtrate CM enhanced dysplastic progression of metaplastic gastroids compared to 50 kDa filtrate. Therefore, they looked specifically for genes encoding 30 kDa or fewer proteins and found high SPON2 expression only in metaplasia and cancer-derived fibroblasts. SPON2 is a 30 kDa secreted protein shown to promote the proliferation and migration of gastric cancer cells. Therefore, they hypothesize that fibroblasts secrete SPON2 to induce dysplasia. The presence of SPON2 in CM was confirmed with mass spectrometry, detecting SPON2 only in metaplasia and cancer-derived fibroblast CM. Furthermore, STAT3 inhibition was evaluated because SPON2 has been shown to promote downstream signaling through STAT3. A STAT3 inhibitor suppressed the effect of cancer-derived fibroblast CM in the dysplastic progression of metaplastic gastroids. These are the first studies investigating fibroblast-derived SPON2 in the induction of precancerous gastric dysplasia. Understanding factors that contribute to gastric dysplasia represents an important area of research, as gastric cancer remains one of the leading causes of cancer-related deaths worldwide.

Yunli (Emily) Chu
(Mentor Brent Ferrell, PhD)
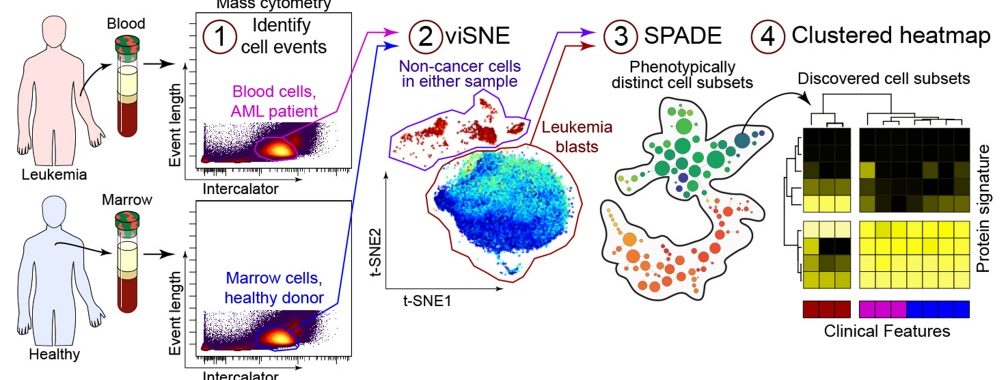
Emily won the Best Talk award at the 23rd Annual Retreat in Cancer Biology! Emily’s research discusses Cytotoxic chemotherapy, which has been the mainstay of acute myeloid leukemia (AML) treatment. There are no approved immunotherapies for AML due to our limited understanding of how CD8+ T cells respond to AML progression. To investigate CD8+ T cell dynamics in AML, we transplanted leukemic cells bearing the MLL-AF9 translocation into immunocompetent mice. Leukemic mice exhibited a shift in CD8+ T cell phenotype toward activation, characterized by reduced naive cells, increased effector cells, and elevated CD69 expression. Surprisingly, high-dimensional phenotyping using mass cytometry identified an enrichment of PD-1 and LAG-3 expressing T cells. Ex vivo stimulation demonstrated heightened TNF-α and IFN-γ production by leukemia-associated T cells. To test if the cytokine milieu is sufficient to induce the phenotypic changes, we stimulated isolated, naive CD8+ T cells in conditioned media containing serum from healthy or leukemic mice. Interestingly, exposure to the AML cytokine milieu alone failed to induce the phenotypic changes observed in vivo. These findings suggest that other factors, such as receptor-ligand interactions and nutrient competition, may play more prominent roles in shaping T-cell responses in AML. In summary, these studies highlight that AML can profoundly influence the phenotype and function of CD8+ T cells. AML induces a dysfunctional phenotype in CD8+ T cells, though functionally, these cells produced more cytokines. It is possible that the secretion of inflammatory cytokines by CD8+ T cells contributes to AML cell proliferation. Future studies will focus on the mechanisms underlying phenotypic changes and immune escape in AML, seeking ways to disrupt these interactions.