Workflow
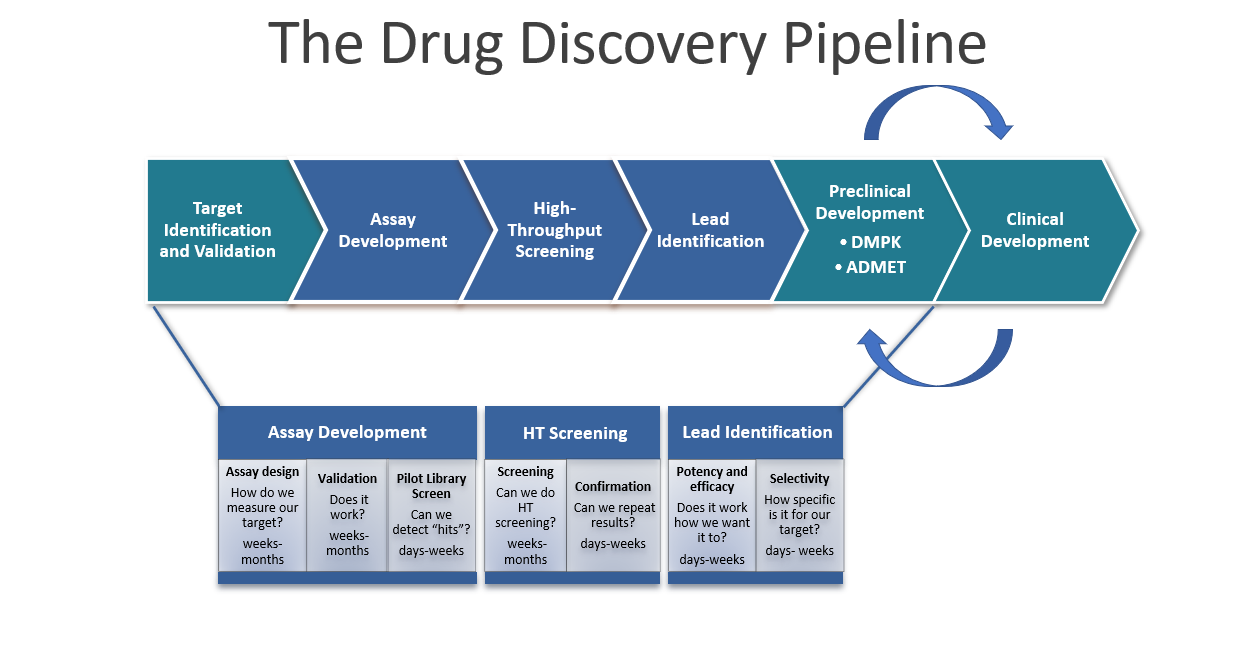
The High-Throughput Screening (HTS) and Molecular Design & Synthesis Center (MDSC) partner with investigators to discover and optimize compounds for chemical probes and drug development.
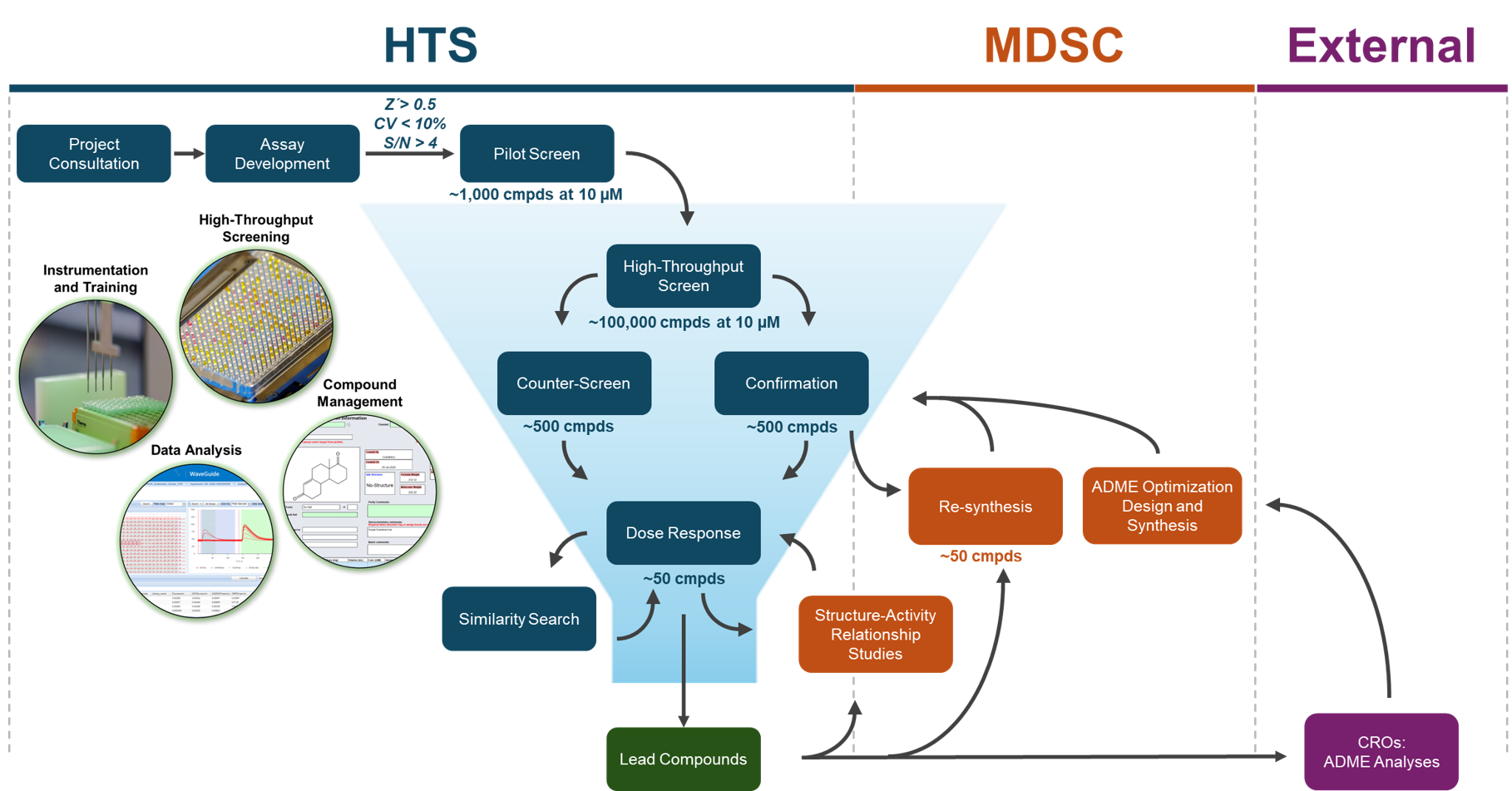
Projects begin with a consultation to set goals, define success measures, and adapt assays for automated HTS. HTS staff assist with assay development, validation, and pilot screening to ensure robust performance. Once optimized, a full screen identifies initial hits (~0.5%), followed by confirmation tests, re-synthesis or vendor reorders, and counter-screens to eliminate false positives.
High-confidence hits undergo dose-response testing to assess potency and efficacy. Informatics-driven similarity searches and SAR studies, supported by MDSC chemists and pharmacologists, further refine compound activity, selectivity, and drug-like properties. For translational projects, HTS/MDSC can coordinate ADME analyses with CROs to inform medicinal chemistry optimization.
HTS staff provide guidance throughout—managing compounds, running screens, analyzing data, and delivering tailored reports. The goal is to generate a focused set of high-quality compounds ready for use as chemical probes or as leads for drug development.
Compound Library
V-HTS Compound Collections
Target-based
Highlighted user ‘success’ stories
Over the past 10 years, the utilization of Panoptic in the Vanderbilt HTS Center, has contributed to more than 50 publications and has supported successful research projects and proposals involving the discovery and characterization of small molecule modulators of ion channels (e.g., Kv, IRKs, SLACK, VRAC, SLO3, etc.), ion transporters (such as KCC2) and GPCRs (e.g., 5-HTRs, mAchRs, mGluRs), the development of novel bioluminescent assay reporters (e.g., CalfluxCTN), and the identification of new activator chemotypes for enzymes (such as NAPE-PLD).
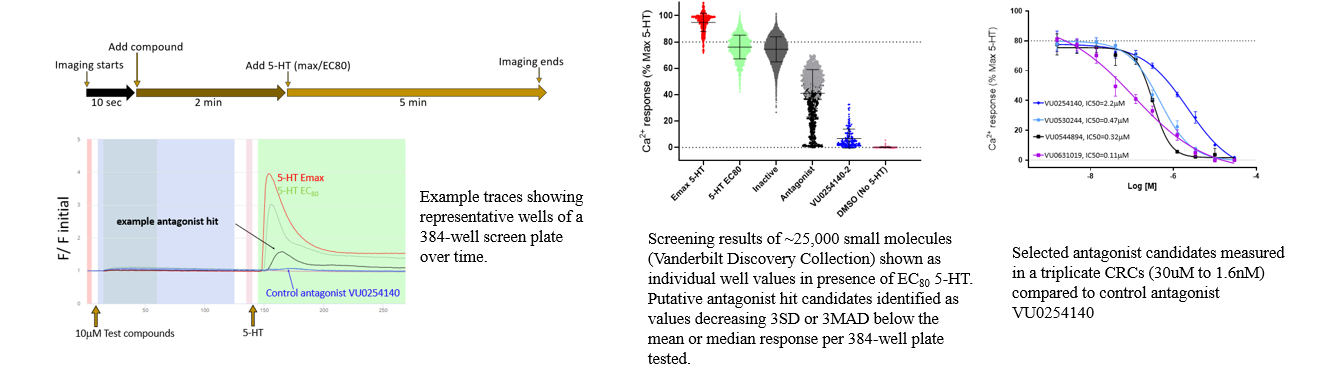
1. Ca2+ Flux Assay: HTS Identifies Potent 5-HT2B Antagonists
Reference: Bender, Aaron M., et al. "Identification of Potent, Selective, and Peripherally Restricted Serotonin Receptor 2B Antagonists from a High-Throughput Screen." ASSAY and Drug Development Technologies 21.3 (2023): 89-96.
Antagonists of the serotonin receptor 2B (5-HT2B) have shown great promise as therapeutics for the treatment of pulmonary arterial hypertension, valvular heart disease, and related cardiopathies. A high-throughput screen performed on Panoptic Imaging Platform led to the identification of highly potent 5-HT2B antagonists (e.g. VU0530244, VU0544894 and VU0631019).
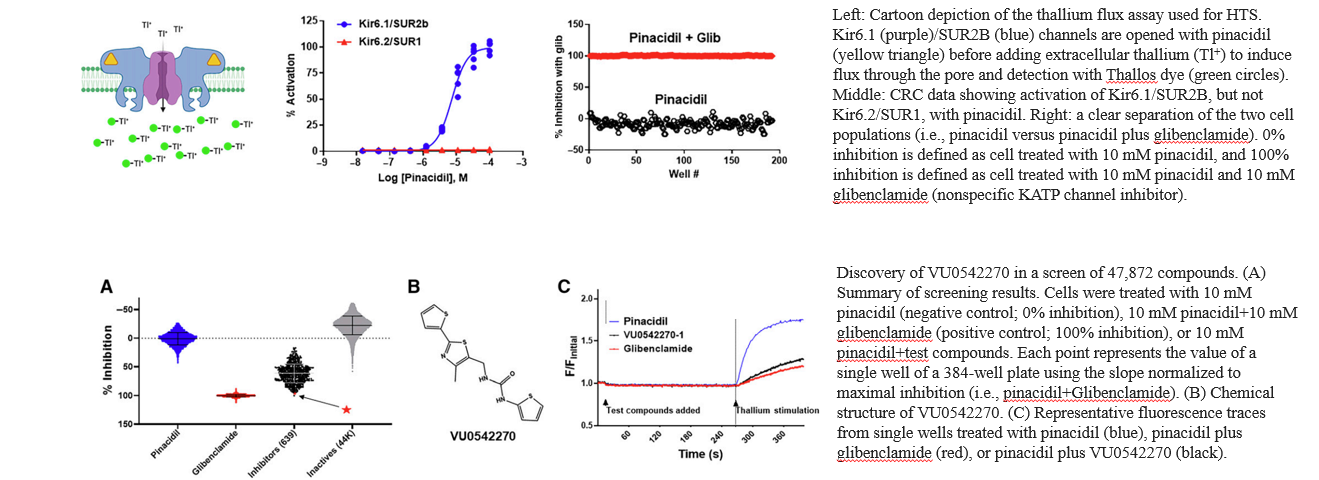
2. Tl+ Flux Assay: HTS Identifies the First Selective Inhibitor or Vascular Kir6.1/SUR2B KATP Channels
Reference: Li, Kangjun, et al. "Discovery and characterization of VU0542270, the first selective inhibitor of vascular Kir6. 1/SUR2B KATP channels." Molecular pharmacology 105.3 (2024): 202-212.
Small-molecule inhibitors of vascular smooth muscle Kir6.1/SUR2B KATP channels represents novel therapeutics for patent ductus arteriosus, migraine headache, and sepsis; however, the lack of selective channel inhibitors has slowed progress in these therapeutic areas. A high-throughput screen performed on Panoptic Imaging Platform led to the discovery of the first vascular-specific KATP channel inhibitor, VU0542270.
Phenotypic (HCI) Screening
coming soon
Functional Genomics Screening
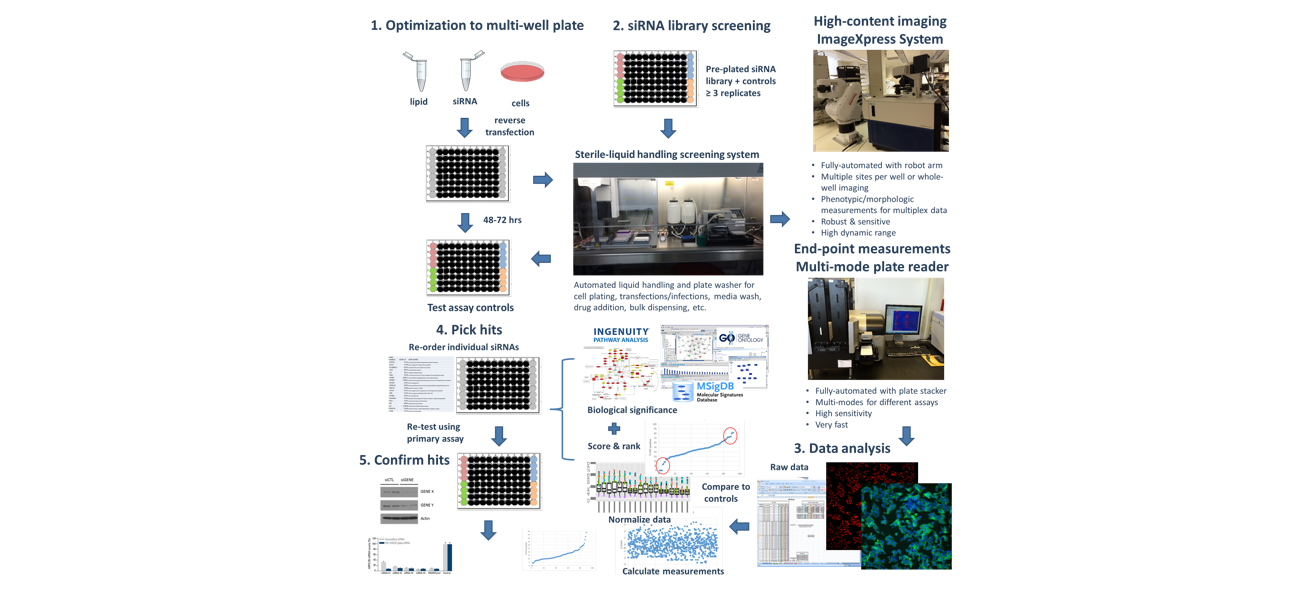
Functional genomics (FG) enables exploration of gene and protein function at the individual gene level as well as the genome-wide scale. RNA interference (RNAi) and gene editing (e.g. CRISPR) alter gene expression/function allowing us to investigate each gene function in a single experiment. The V-HTS Functional Genomics Screening service provides tools, technologies, protocols, and access for individual genes, pathway-focused genes, and genome-wide screening capabilities. V-HTS has standardized most protocols for FGS services and has expertise in customized screens to meet your needs. The core serves as a forum for scientific exchange and nurturing collaborations in FGS at Vanderbilt and research community.
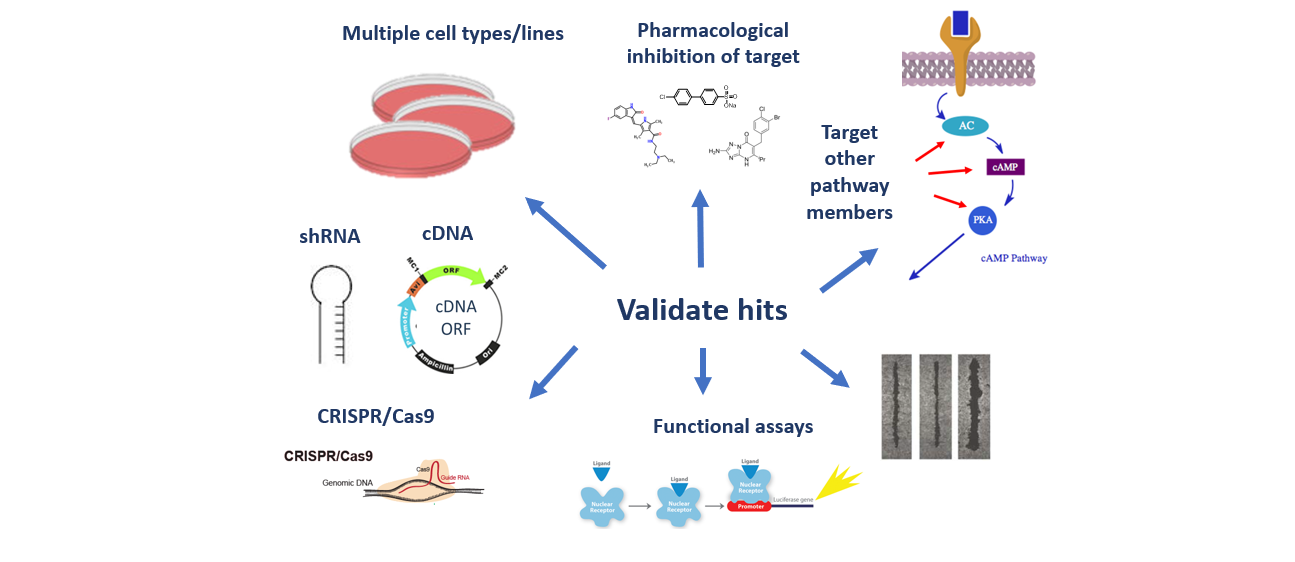
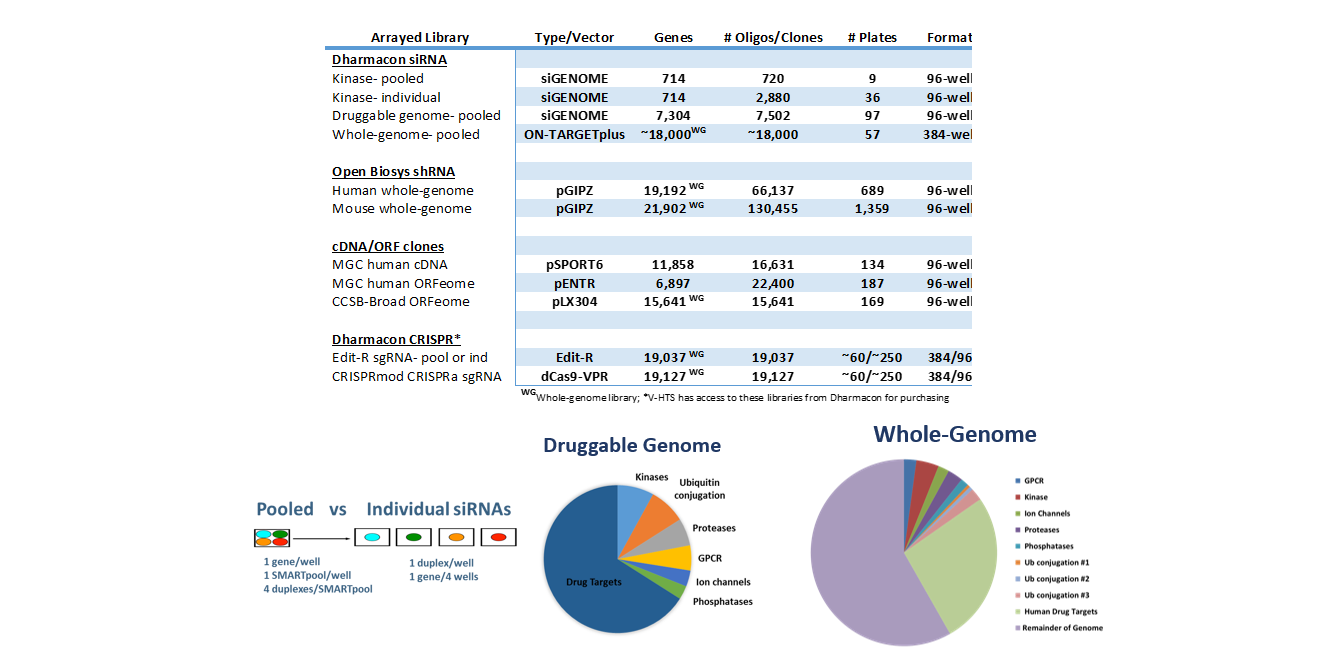
V-HTS Functional Genomics Screening Libraries
Timeline
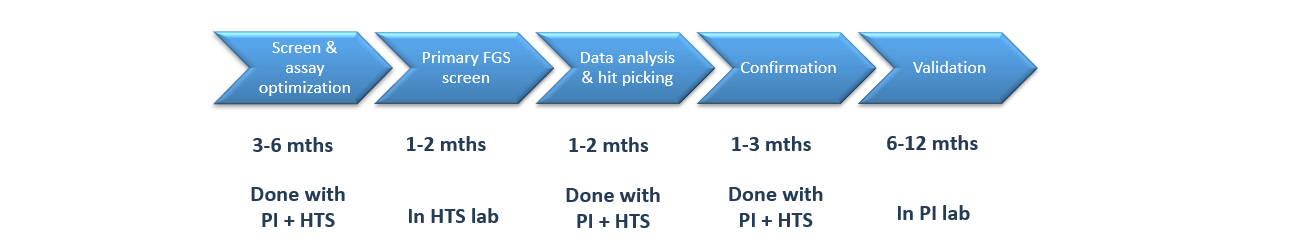
Full-Service Workflow