By Lorena Infante Lara
Food-related allergies and their effects are insidiously common in the United States: over seven percent of children and over 10 percent of adults are affected. Although a quarter of people with food allergies have an epinephrine pen, they are expensive and have to be replaced about once a year due to short expiration dates. About 40 percent of people with food allergies have at least one life-threatening, food allergy–related emergency department visit in their lifetimes.
But exactly what is the leading, food allergy–related cause of death in the U.S. and the United Kingdom? Peanut and tree nut allergies.
Despite its prevalence, the molecular details of how peanut allergies are induced and expressed in the body are not clear. A recent pair of papers from the Vanderbilt University labs of Benjamin Spiller, associate professor of pharmacology, and Scott Smith, associate professor of medicine, dig into how peanut allergies are provoked and providing support for the use of a potential treatment option: hypoallergens. Both papers were published in February in the Journal of Allergy and Clinical Immunology.
IgE antibodies, which are responsible for allergies, are also the least abundant antibody type in the bloodstream and have been difficult to study. To tackle this problem, Smith and colleagues created hybrid cell lines that reliably secreted IgE antibodies against peanuts. This allowed them to test the antibodies’ structural, immunological, and functional properties and how they interact with peanut allergens.
Thanks to a comprehensive analysis, the first paper, “Antigenic determinants underlying IgE-mediated anaphylaxis to peanut,” shows that the antibodies bind most commonly to five regions within two peanut proteins. Collaborator Stokes Peebles, Elizabeth and John Murray Professor of Medicine, verified these results in an animal model, revealing which combination of antibodies binding at which protein regions can cause a life-threatening allergic reaction. Additional testing in patients with a peanut allergy validated which specific regions result in the worst allergic reactions.
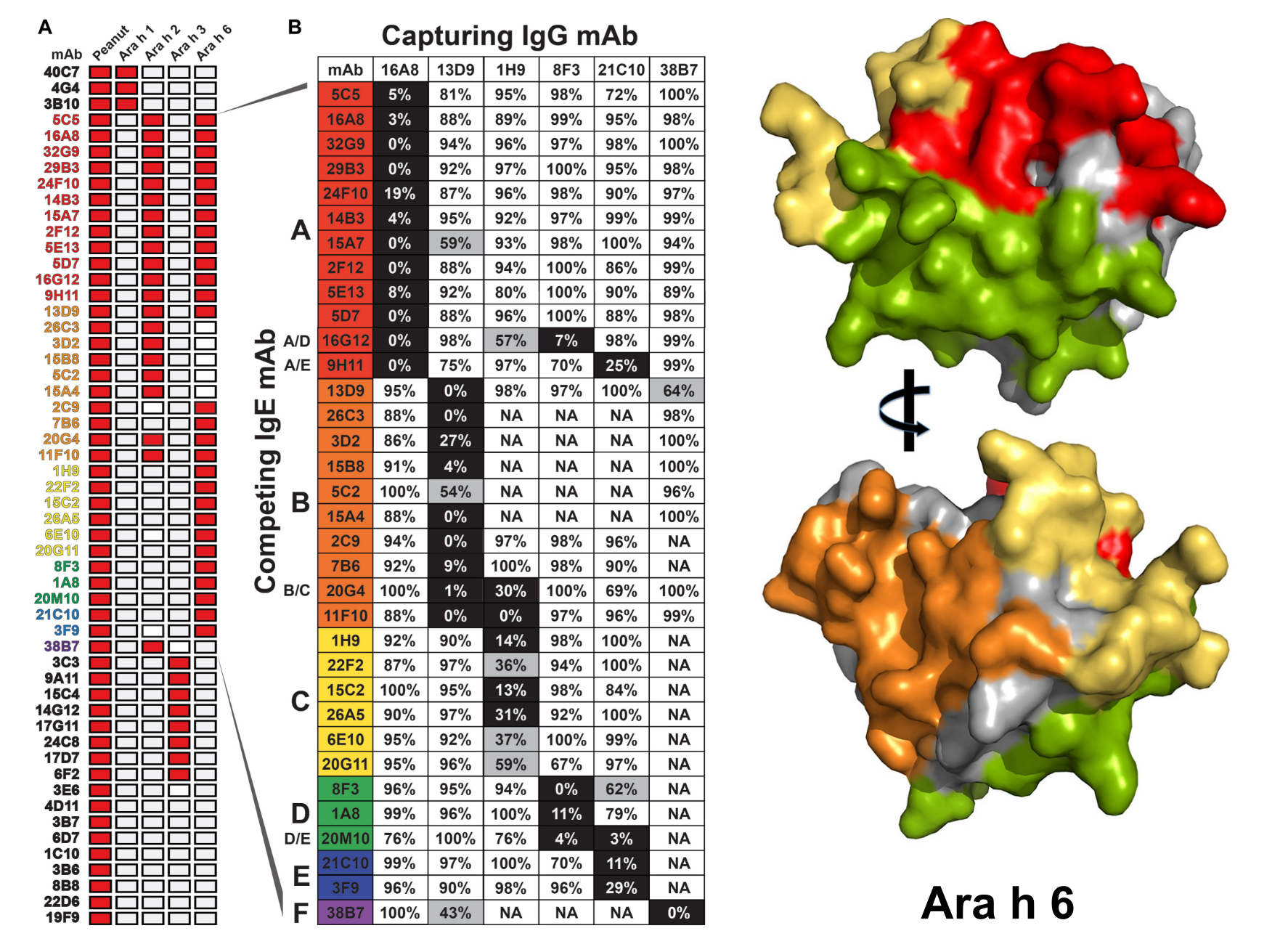
“Diagnostic tests that capture the fine detail of the antibodies responsible for disease in a given patient will be essential … to identify instances where other peanut allergens are important for disease” and to guide an individual’s treatment as new therapeutic strategies emerge, the researchers said.
In their second paper, “Structural determinants of peanut induced anaphylaxis,” the Vanderbilt biomedical researchers visualized, in vast detail, the physical interfaces between the antibodies and their allergens.
“Now knowing how the antibodies interacted with the antigens, we generated antibody variants that disturbed that interaction and found that the impaired binding did not lead to anaphylaxis,” Spiller said. “These results validate ‘hypoallergens’ as a potential therapeutic against peanut allergy, which we hope to use as vaccines in sensitive individuals.”
A hypoallergen treatment would expose a person to modified peanut proteins that elicit lesser immune reactions. This treatment would reduce the strength of the allergic reaction, making the desensitization process easier.
Until we can stem the tide of food allergy incidence, our best bet is to treat it, and hypoallergens are a promising tactic. Basic molecular and structural research on the inner workings of allergy, such as the one presented here, will help to inform clinicians’ and scientists’ efforts to understand and treat food allergies.
Go deeper
The paper “Antigenic determinants underlying IgE-mediated anaphylaxis to peanut” was published in the Journal of Allergy and Clinical Immunology in February 2025. Its companion paper, “Structural determinants of peanut induced anaphylaxis,” was published in the same issue.
Funding
This research used funds from the National Institutes of Health, specifically the National Institute of Allergy and Infectious Disease; Food Allergy Research and Education; a Vanderbilt University Discovery Grant; and a Vanderbilt University Stanley Cohen Innovation Fund award.
Shared resources
This research made use of the Mass Spectrometry Research Center’s Proteomics Core and the Center for Structural Biology’s Computational Structural Biology core and Biophysical Instrumentation core.
